Vignette: Write & Read Multiple Excel files with purrr
Introduction
This post will show you how to write and read a list of data tables to and from Excel with purrr, the functional programming package 📦 from tidyverse. In this example I will also use the packages readxl and writexl for reading and writing in Excel files, and cover methods for both XLSX and CSV (not strictly Excel, but might as well!) files.


Whilst the internet is certainly in no shortage of R tutorials on how to read and write Excel files (see this Stack Overflow thread for example), I think a purrr approach still isn’t as well-known or well-documented. I find this approach to be very clean and readable, and certainly more “tidyverse-consistent” than other approaches which rely on lapply() or for loops. My choice of packages 📦 for reading and writing Excel files are readxl and writexl, where the advantage is that neither of them require external dependencies.
For reading and writing CSV files, I personally have switched back and forth between readr and data.table, depending on whether I have a need to do a particular analysis in data.table (see this discussion on why I sometimes use it in favour of dplyr). Where applicable in this post, I will point out in places where you can use alternatives from data.table for fast reading/writing.
For documentation/demonstration purposes, I’ll make the package references (indicated by ::) explicit in the functions below, but it’s advisable to remove them in “real life” to avoid code that is overly verbose.
Getting Started
The key functions used in this vignette come from three packages: purrr, readxl, and writexl.
library(tidyverse)
library(readxl)
library(writexl)Since purrr is part of core tidyverse, we can simply run library(tidyverse). This is also convenient as we’ll also use various functions such as group_split() from dplyr and the %>% operator from magrittr in the example.
Note that although readxl is part of tidyverse, you’ll still need to load it explicitly as it’s not a “core” tidyverse package.
Writing multiple Excel files
Let us start off with the iris dataset that is pre-loaded with R. If you’re not one of us sad people who almost know this dataset by heart, here’s what it looks like:
iris %>% head()## Sepal.Length Sepal.Width Petal.Length Petal.Width Species
## 1 5.1 3.5 1.4 0.2 setosa
## 2 4.9 3.0 1.4 0.2 setosa
## 3 4.7 3.2 1.3 0.2 setosa
## 4 4.6 3.1 1.5 0.2 setosa
## 5 5.0 3.6 1.4 0.2 setosa
## 6 5.4 3.9 1.7 0.4 setosaThe first thing that we want to do is to create multiple datasets, which we can do so by splitting iris. I’ll do this by running group_split() on the Species column, so that each species of iris has its own dataset. This will return a list of three data frames, one for each unique value in Species: setosa, versicolor, and virginica. I’ll assign these three data frames to a list object called list_of_dfs:
# Split: one data frame per Species
iris %>%
dplyr::group_split(Species) -> list_of_dfs
list_of_dfs## [[1]]
## # A tibble: 50 x 5
## Sepal.Length Sepal.Width Petal.Length Petal.Width Species
## <dbl> <dbl> <dbl> <dbl> <fct>
## 1 5.1 3.5 1.4 0.2 setosa
## 2 4.9 3 1.4 0.2 setosa
## 3 4.7 3.2 1.3 0.2 setosa
## 4 4.6 3.1 1.5 0.2 setosa
## 5 5 3.6 1.4 0.2 setosa
## 6 5.4 3.9 1.7 0.4 setosa
## 7 4.6 3.4 1.4 0.3 setosa
## 8 5 3.4 1.5 0.2 setosa
## 9 4.4 2.9 1.4 0.2 setosa
## 10 4.9 3.1 1.5 0.1 setosa
## # ... with 40 more rows
##
## [[2]]
## # A tibble: 50 x 5
## Sepal.Length Sepal.Width Petal.Length Petal.Width Species
## <dbl> <dbl> <dbl> <dbl> <fct>
## 1 7 3.2 4.7 1.4 versicolor
## 2 6.4 3.2 4.5 1.5 versicolor
## 3 6.9 3.1 4.9 1.5 versicolor
## 4 5.5 2.3 4 1.3 versicolor
## 5 6.5 2.8 4.6 1.5 versicolor
## 6 5.7 2.8 4.5 1.3 versicolor
## 7 6.3 3.3 4.7 1.6 versicolor
## 8 4.9 2.4 3.3 1 versicolor
## 9 6.6 2.9 4.6 1.3 versicolor
## 10 5.2 2.7 3.9 1.4 versicolor
## # ... with 40 more rows
##
## [[3]]
## # A tibble: 50 x 5
## Sepal.Length Sepal.Width Petal.Length Petal.Width Species
## <dbl> <dbl> <dbl> <dbl> <fct>
## 1 6.3 3.3 6 2.5 virginica
## 2 5.8 2.7 5.1 1.9 virginica
## 3 7.1 3 5.9 2.1 virginica
## 4 6.3 2.9 5.6 1.8 virginica
## 5 6.5 3 5.8 2.2 virginica
## 6 7.6 3 6.6 2.1 virginica
## 7 4.9 2.5 4.5 1.7 virginica
## 8 7.3 2.9 6.3 1.8 virginica
## 9 6.7 2.5 5.8 1.8 virginica
## 10 7.2 3.6 6.1 2.5 virginica
## # ... with 40 more rowsI’ll also use purrr::map() to take the character values (setosa, versicolor, and virginica) from the Species column itself for assigning names to the list. map() transforms an input by applying a function to each element of the input, and then returns a vector the same length as the input. In this immediate example, the input is the list_of_dfs and we apply the function dplyr::pull() to extract the Species variable from each data frame. We then repeat this approach to convert Species into character type with as.character() and take out a single value with unique():
# Use the value from the "Species" column to provide a name for the list members
list_of_dfs %>%
purrr::map(~pull(.,Species)) %>% # Pull out Species variable
purrr::map(~as.character(.)) %>% # Convert factor to character
purrr::map(~unique(.)) -> names(list_of_dfs) # Set this as names for list members
names(list_of_dfs)## [1] "setosa" "versicolor" "virginica"These names will be useful for exporting the data frames into Excel, as they will effectively be our Excel sheet names. You can always manually hard-code the sheet names, but the above approach allows you to do the entire thing dynamically if you need to.
Having set the sheet names, I can then pipe the list of data frames directly into write_xlsx(), where the Excel file name and path is specified in the same path argument:
list_of_dfs %>%
writexl::write_xlsx(path = "../datasets/test-excel/test-excel.xlsx")Writing multiple CSV files
Exporting the list of data frames into multiple CSV files will take a few more lines of code, but still relatively straightforward. There are three main steps:
Define a function that tells R what the names for each CSV file should be, which I’ve called
output_csv()below. The data argument will take in a data frame whilst the names argument will take in a character string that will form part of the file name for the individual CSV file.Create a named list where the names match the arguments of the function you’ve just defined (data and names), and should contain the objects that you would like to pass through to the function for the respective arguments. In this case, list_of_dfs will provide the three data frames, and names(list_of_dfs) will provide the names of those three data frames. This is necessary for running
pmap(), which in my view is basically a super-powered version ofmap()that lets you iterate over multiple inputs simultaneously.pmap()will then iterate through the two sets of inputs throughoutput_csv()(the inputs are used as arguments), which then writes the three CSV files with the file names you want. For the “writing” function, you could either usewrite_csv()from readr (part of tidyverse) orfwrite()from data.table, depending on your workflow / style.
# Step 1
# Define a function for exporting csv with the desired file names and into the right path
output_csv <- function(data, names){
folder_path <- "../datasets/test-excel/"
write_csv(data, paste0(folder_path, "test-excel-", names, ".csv"))
}
# Step 2
list(data = list_of_dfs,
names = names(list_of_dfs)) %>%
# Step 3
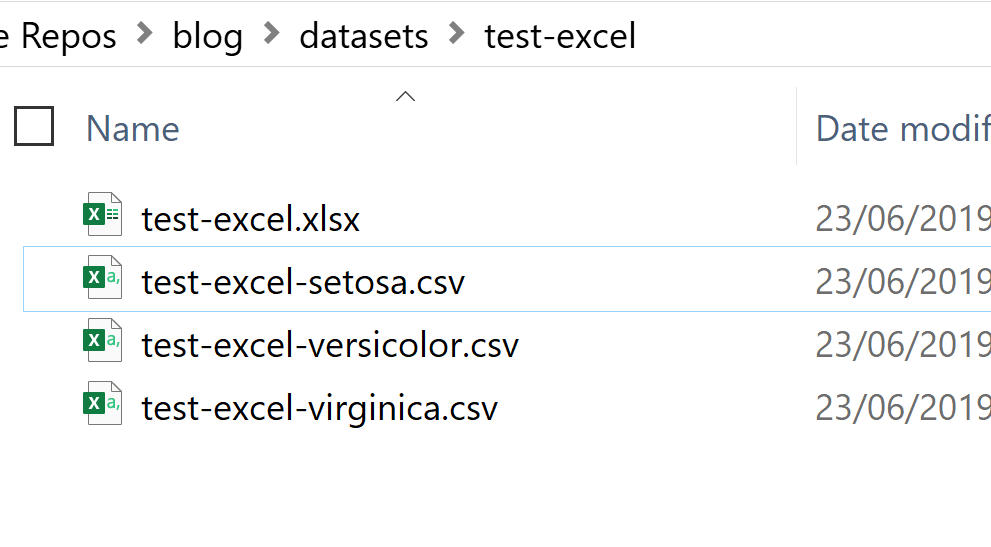
purrr::pmap(output_csv) The outcome of the above code is shown below. My directory now contains one Excel file with three Worksheets (sheet names are “setosa”, “versicolor”, and “virginica”), and three separate CSV files for each data slice:
Reading multiple Excel / CSV files

For reading files in, you’ll need to decide on how you want them to be read in. The options are:
- Read all the datasets directly into the Global Environment as individual data frames with a “separate existence” and separate names.
- Read all the datasets into a single list, where each data frame is a member of that list.
The first option is best if you are unlikely to run similar operations on all the data frames at the same time. You may for instance want to do this if the data sets that you are reading in are structurally different from each other, and that you are planning to manipulate them separately.
The second option will be best if you are likely to manipulate all the data frames at the same time, where for instance you may run on the list of data frames map() with drop_na() as an argument to remove missing values for all of the data frames at the same time. The benefit of reading your multiple data sets into a list is that you will have a much cleaner workspace (Global Environment). However, there is a minor and almost negligible inconvenience accessing individual data frames, as you will need to go into a list and pick out the right member of the list (e.g. doing something like list_of_dfs[3]).
Method 1A: Read all sheets in Excel into Global Environment
So let’s begin! This method will read all the sheets within a specified Excel file and load them into the Global Environment, using variable names of your own choice. For simplicity, I will use the original Excel sheet names as the variable names.
The first thing to do is to specify the file path to the Excel file:
wb_source <- "../datasets/test-excel/test-excel.xlsx"You can then run readxl::excel_sheets() to extract the sheet names in that Excel file, and save it as a character type vector.
# Extract the sheet names as a character string vector
wb_sheets <- readxl::excel_sheets(wb_source)
print(wb_sheets)## [1] "setosa" "versicolor" "virginica"The next step is to iterate through the sheet names (saved in wb_sheets) using map(), and within each iteration use assign() (base) and read_xlsx() (from readxl) to load each individual sheet into the Global Environment, giving each one a variable name. Here’s the code:
# Load everything into the Global Environment
wb_sheets %>%
purrr::map(function(sheet){ # iterate through each sheet name
assign(x = sheet,
value = readxl::read_xlsx(path = wb_source, sheet = sheet),
envir = .GlobalEnv)
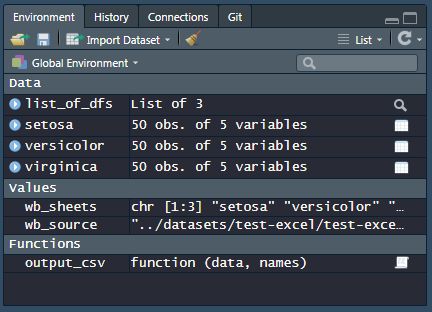
})This is what my work space looks like:
Note that map() always returns a list, but in this case we do not need a list returned and only require the “side effects”, i.e. the objects being read in to be assigned to the Global Environment. If you prefer you can use lapply() instead of map(), which for this purpose doesn’t make a big practical difference.
Also, assign() allows you to assign a value to a name in an environment, where we’ve specified the following as arguments:
- x:
sheetas the variable name - value: The actual data from the sheet we read in. Here, we use
readxl::read_xlsx()for reading in specific sheets from the Excel file, where you simply specify the file path and the sheet name as the arguments. - envir:
.GlobalEnvas the environment
Method 1B: Read all CSV files in directory into Global Environment
The method for reading CSV files into a directory is slightly different, as you’ll need to find a way to identify or create a character vector of names of all the files that you want to load into R. To do this, we’ll use list.files(), which produces a character vector of the names of files or directories in the named directory:
file_path <- "../datasets/test-excel/"
file_path %>% list.files()## [1] "test-excel-setosa.csv" "test-excel-versicolor.csv"
## [3] "test-excel-virginica.csv" "test-excel.xlsx"We only want CSV files in this instance, so we’ll want to do a bit of string manipulation (using str_detect() from stringr - again, from tidyverse) to get only the names that end with the extension “.csv”. Let’s pipe this along:
file_path %>%
list.files() %>%
.[str_detect(., ".csv")] -> csv_file_names
csv_file_names## [1] "test-excel-setosa.csv" "test-excel-versicolor.csv"
## [3] "test-excel-virginica.csv"The next part is similar to what we’ve done earlier, using map(). Note that apart from replacing the value argument with read_csv() (or you can use fread() to return a data.table object rather than a tibble), I also removed the file extension in the x argument so that the variable names would not contain the actual characters “.csv”:
# Load everything into the Global Environment
csv_file_names %>%
purrr::map(function(file_name){ # iterate through each file name
assign(x = str_remove(file_name, ".csv"), # Remove file extension ".csv"
value = read_csv(paste0(file_path, file_name)),
envir = .GlobalEnv)
})Method 2A: Read all sheets in Excel into a list
Reading sheets into a list is actually easier than to read it into the Global Environment, as map() returns a list and you won’t have to use assign() or specify a variable name. Recall that wb_source holds the path of the Excel file, and wb_sheets is a character vector of all the sheet names in the Excel file:
# Load everything into the Global Environment
wb_sheets %>%
purrr::map(function(sheet){ # iterate through each sheet name
readxl::read_xlsx(path = wb_source, sheet = sheet)
}) -> df_list_read # Assign to a listYou can then use map() again to run operations across all members of the list, and even chain operations within it:
df_list_read %>%
map(~select(., Petal.Length, Species) %>%
head())## [[1]]
## # A tibble: 6 x 2
## Petal.Length Species
## <dbl> <chr>
## 1 1.4 setosa
## 2 1.4 setosa
## 3 1.3 setosa
## 4 1.5 setosa
## 5 1.4 setosa
## 6 1.7 setosa
##
## [[2]]
## # A tibble: 6 x 2
## Petal.Length Species
## <dbl> <chr>
## 1 4.7 versicolor
## 2 4.5 versicolor
## 3 4.9 versicolor
## 4 4 versicolor
## 5 4.6 versicolor
## 6 4.5 versicolor
##
## [[3]]
## # A tibble: 6 x 2
## Petal.Length Species
## <dbl> <chr>
## 1 6 virginica
## 2 5.1 virginica
## 3 5.9 virginica
## 4 5.6 virginica
## 5 5.8 virginica
## 6 6.6 virginicaMethod 2B: Read all CSV files in directory into a list
At this point you’ve probably gathered how you can adapt the code to read CSV files into a list, but let’s cover this for comprehensiveness. No assign() needed, and only run read_csv() within the map() function, iterating through the file names:
# Load everything into the Global Environment
csv_file_names %>%
purrr::map(function(file_name){ # iterate through each file name
read_csv(paste0(file_path, file_name))
}) -> df_list_read2 # Assign to a listThank you for reading! 😄
Hopefully this is a helpful tutorial for an iterative approach to writing and reading Excel files. If you like what you read or if you have any suggestions / thoughts about the subject, do leave a comment in the Disqus fields in the blog and let me know!